Quick-Compute Tools
Online calculators and tools for quick and easy access to computational neuroscience methods.
k-mer generator
Our genetic analysis services provide insights into the genetic factors influencing Parkinson's disease and brain health.
Extrapolation of total counts, region volume, region area, and density from systematic random sampling of a larger area or volume.
Stereology Cell Counts
Data Tables
Parkinson's Disease Cell Markers
Research for Parkinson's disease hinges on characterization of degenerative changes in specific cell types. To aid researchers and clinicians in understanding the complexities of this condition, a specialized table of cell markers has been developed. This lookup table allows users to categorize and define various groups of central nervous system cells based on genetic markers.
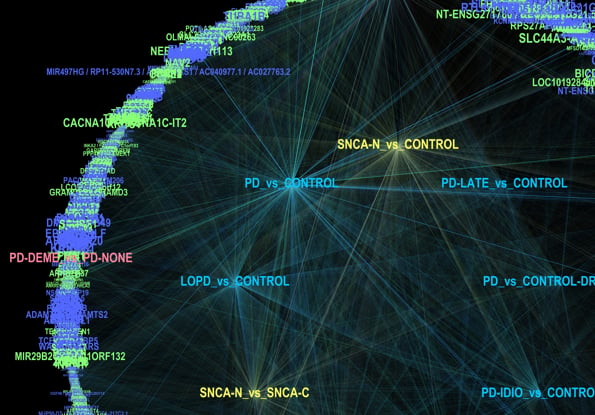
Parkinson's Disease -omics Human Research Findings
This repository represents a collection of datasets from main text and supplemental tables presenting findings from various next-generation sequencing and proteomic studies. Careful collation of these datasets, allows for a comparison of transcriptomic and proteomic markers across different research projects. This compilation facilitates the identification of correlations and discrepancies across studies.
Parkinson's Disease Brain Pathology
Coming soon!
We are in the process of developing a database of pathological findings from human brain tissue studies. This database will serve as a critical resource for understanding progression at different brain sites. Our goal is to ensure that the database is user-friendly and easily accessible, thus promoting collaboration across the scientific community.
ATAC-seq:RNA-seq Plots
Code
Make plots for ATAC-seq and RNA-seq preliminary viewing. These are not final plots that account for distance and do not incorporate gtf coordinates.
Input: Differential expression and differential accessibility data
Output: Bar plots in display and as saved images
Retrieve Data Ensembl
PubMed Output Processing
Clustering from DAVID
Will define best cluster number and make plots for DAVID gene annotation outputs.
Input: DAVID Functional Annotation Chart tables in text format
Output: Silhouette plots and scatter plots made with principal component analysis and t-distributed stochastic neighbor embedding in display and as saved images
Takes PubMed input that contains metainformation associated with publications.
Input: PubMed-output text files from literature searches
Output: Two text files, one with all fields and information, and one with specified fields
Looks up genes in the most recent Ensembl update and extracts information needed for matching IDs across studies
Input: List of gene symbols or ENSG ids
Output: Files with pertinent information
Agglomerative Hierarchical Clustering
Clusters data by various distance methods
Input: Data matrix of values
Output: Dendrograms in viewer
Code used and included on Tools page that accounts for sampling fractions.
Input: User defined through HTML form
Output: Calculation in HTML viewer
Counts Calculator
k-mer Generator
Takes a longer sequence and cuts it up, outputting a fraction of possibilities
Input: User input in form
Output: Text list of k-mers
Heatmap
Takes data from any type of study and makes heatmaps of values.
Input: Data matrix of values
Output: Heatmap in viewer and as saved file
Principal Component Analysis
Dimensional reduction by principal component analysis, making two hypothetical factors each for variables and subjects.
Input: Data matrix of values
Output: Plots and text information on analysis in viewer
k-mer Generator
Code used and included on Tools page that accounts for sampling fractions.
Input: User defined through HTML form
Output: Calculation in HTML viewer